To better understand how your variants might be linked to disease, you can add information from DisGeNET to your VEP analysis. DisGeNET integrates data from expert curated repositories, GWAS catalogues, animal models and the scientific literature, to provide disease associations with genes and variants.
Continue readingAuthor: Emily (Outreach)
Cool stuff Ensembl VEP can do: links to variation databases
When you find a variant you are interested in, you will want to thoroughly investigate all current knowledge about it. For each Ensembl release we update our variation databases with the latest public information from a wide range of resources and make summary information available in VEP. We now make it easier to link to fuller details in these resources by providing lists of names used for each variant in each.
Continue readingCool stuff the VEP can do: publication data from Mastermind
With absolutely tons of publication data out there, finding relevant information about your variants and genes of interest can be a struggle. We’ve been working with Genomenon to integrate Mastermind, the Genomic Search Engine, into VEP results, bringing you publication data about genes, variants, phenotypes, diseases and therapy evidence.
Continue readingEnsembl 103 has been released
Squeak squeak, we’ve got a new release out! In Ensembl 103 we’ve got the latest mouse genome assembly, updates to human genes, variation and regulation, new and updated genomes for vertebrates, metazoa and plants and a new tool for converting variant descriptions.
Continue readingUpdates to the Ensembl Variant Recoder
The Variant Recoder, available on the Ensembl REST API, can help you with data re-use. Multiple identifiers, coming from different databases, can refer to the same variant – the Variant Recoder can help.
Continue readingWhat’s coming in release 103
We’re planning to release Ensembl 103 in February 2021. We’ve got lots of plans, although we can’t guarantee everything will make it into the release.
Continue readingMANE progress update
NCBI and EBI have been hard at work on our joint MANE collaboration, aiming to provide a set of representative transcripts for human protein-coding genes that are identically annotated in the NCBI RefSeq and Ensembl/GENCODE annotation sets and exactly match the GRCh38 reference assembly. We released MANE v0.5 in Dec 2018, which included one well-supported MANE Select transcript for 53% of the human protein-coding genes. The remainder has required a lot more analysis and curation than we expected, but we’re pleased to announce MANE v0.92, now covering 16,865 genes or ~88% of known protein-coding genes. We’ve been focussing on clinically relevant genes and MANE Select now includes 99% of genes with high gene-disease validity. This release also includes 43 extra transcripts labeled “MANE Plus Clinical” that have been chosen to aid in clinical reporting, for example when there are additional pathogenic or likely pathogenic variants not covered in the MANE Select transcript. For example in genes where there are mutually exclusive exons and both exons have clinically relevant variants, a MANE Plus Clinical transcript will be added alongside the MANE Select transcript so that both exons are represented in MANE.
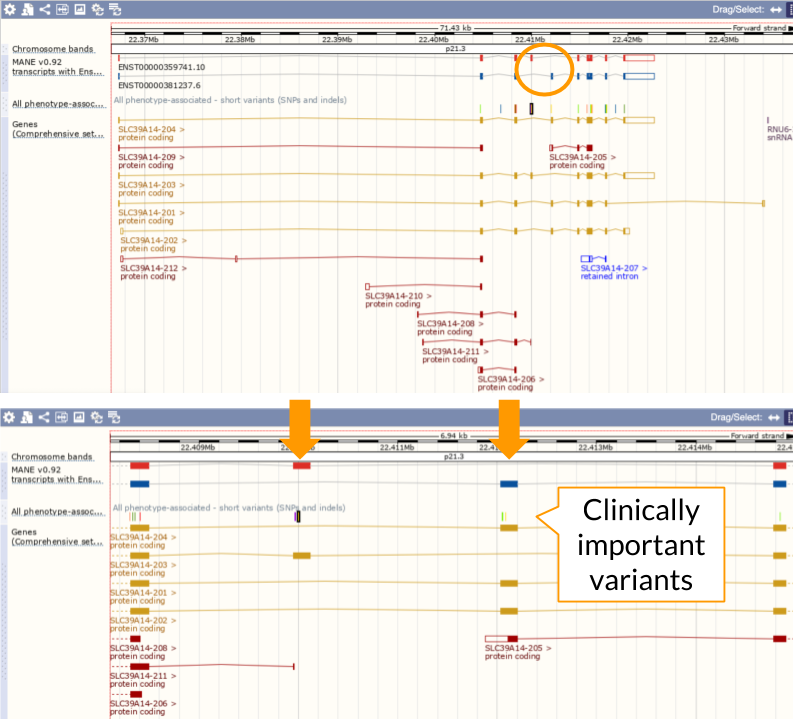
While it’s critical to consider other alternatively-spliced transcripts for variant interpretation or functional analyses, the MANE Select and MANE Plus Clinical transcripts provide a common foundation for clinical reporting, and other analyses that benefit from using just one well-supported transcript or protein per gene.
MANE Select is now shown in the genome aggregation database gnomAD v3, is displayed and used as the preferred transcript for variant reporting in ClinVar and is displayed in DECIPHER. We have released this data as a trackhub for display in the Ensembl, NCBI and UCSC genome browsers. MANE Select v0.92 transcripts will be available in Ensembl release 103 due in the Spring 2021, and will be included in BioMart and VEP.
Partnership with the community is really important to us. We value your feedback. If you are interested in working with us, please contact us at MANE-help@ebi.ac.uk.
Cool stuff Ensembl VEP can do: one click BioMart
The web VEP tool gives you a lot of useful information about your variants, but you may want more details, for example about the genes your variants overlap: maybe you want to fetch their sequences, homologues or protein domains. With a single click, you can go straight to BioMart, using your list of genes or known variants as your filter.
Continue readingGene and protein families retiring
As of release 102, we will be removing the gene and protein family analysis from Ensembl. Information on related genes within and between species is still provided by our gene trees, homologues and alignments.
Continue readingCool stuff Ensembl VEP can do – What’s in the cache and how does VEP use it?
We often get questions about Ensembl VEP caches – the compressed data files we create for each new Ensembl release, which can be automatically installed with the VEP installer script – so here’s a quick introduction to these handy data bundles.
Continue reading