Ensembl 91 is now live! The cat is truly out of the bag now, and we can safely say that there’s been no monkeying around this release; we’ve been very busy!
Read on to discover the highlights of this new release and be sure to join us for our release webinar at 16:00 (GMT) on Wednesday 13th December for a guided tour of the new data and site.
New species, gene sets and annotations
Twelve new Primate Genomes:
- Nancy Ma’s night monkey (Aotus nancymaae)
- White-headed capuchin (Cebus capucinus)
- Sooty mangabey (Cercocebus atys)
- Angola colobus (Colobus angolensis ssp. palliatus)
- Crab-eating macaque (Macaca fasicularis)
- Southern pig-tailed macaque (Macaca nemestrina)
- Drill (Mandrillus leucophaeus)
- Bonobo (Pan paniscus)
- Coquerel’s sifaka (Propithecus coquereli)
- Black snub-nosed monkey (Rhinopithecus bieti)
- Golden snub-nosed monkey (Rhinopithecus roxellana)
- Black-capped/Bolivian squirrel monkey (Saimiri boliviensis ssp. boliviensis)

The delightful new faces of Ensembl – the twelve new primate species added for Release 91
Updates to six existing primate genomes:
- Chimpanzee (Pan troglodytes, Pan_tro_3.0)
- Gibbon (Nomascus leucogenys, Nleu_3.0)
- Gorilla (Gorilla gorilla spp. gorilla, gorGor4)
- Mouse lemur (Microcebus murinus, Mmur_3.0)
- Olive baboon (Papio anubis, Panu_3.0)
- Tarsier (Carlito syrichta, Tarsius_syrichta-2.0.1)
All comparative genomics tools and views will now include these species, and microarray probe mappings have been added.
Updates to other species:
Cinnamon the cat is back in the spotlight as we have also incorporated and annotated the new Cat genome assembly, Felis catus v8.0. The Ensembl-HAVANA GENCODE gene set has also been updated for mouse (vM16), and both human and mouse species also have new cDNA databases. For the mouse genome, the latest (v21) Consensus Coding Sequence (CCDS) dataset has also been imported. We also have updated annotation for Fruitfly (dmel_r6.17) from the FlyBase release FB2017_04 and C. elegans from WormBase release WS260.
New tool: Linkage Disequilibrium Calculator for Variation data
Are you interested in finding out whether your variants of interest are linked, or commonly inherited together? We have produced a new tool where you can calculate linkage disequilibrium (LD) between variants. Currently available for human data only, you can find LD data for all the different populations in the 1000 Genomes Project. You can provide a genomic region of interest (specifying genomic coordinates) or with a list of variants. You can also define the ‘window size’ where you can investigate variants unto 250,000 base pairs upstream and downstream (maximum total window size of 500,000bp) of your variant of interest.
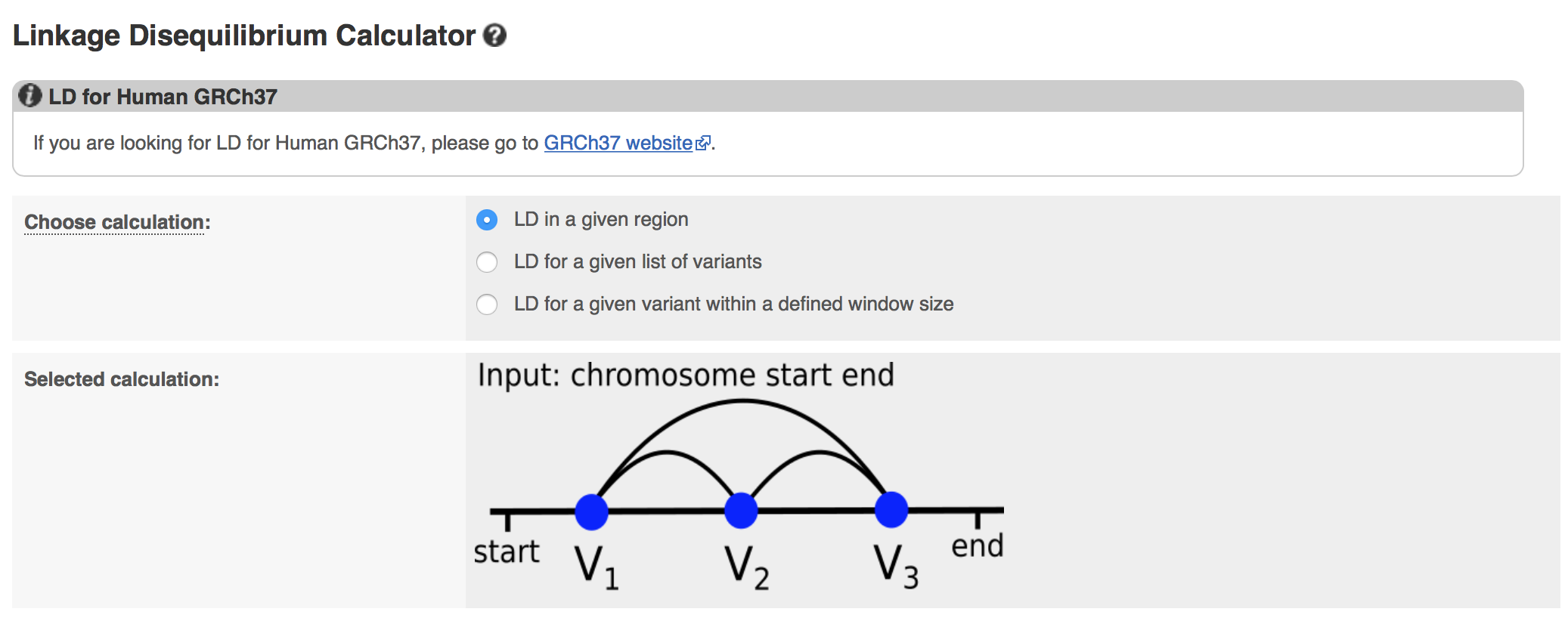
A screenshot of our new Linkage Disequilibrium tool
Variation Data Updates
Updated and novel variation data for human:
- Cancer variation data has been incorporated from version 82 of the Catalogue of Somatic Mutations in Cancer (COSMIC).
- Updates to the Human Gene Mutation Database (HGMD) data to version 2017.2.
- Structural variation data from the Database of Genomic Variants archive (DGVa). New study data incorporated and updates to existing structural data.
- Direct links to PharmGKB on variant pages, so you can quickly discover the effect of a variant on drug response.
- Phenotype data reviewed and updates. Data has been incorporated from:
- NHGRI-EBI GWAS catalog
- OMIM and MIM morbid
- ClinVar
- UniProt
- COSMIC Gene Census
- DDG2p
- Orphanet
Other species variation data updates include:
- New dbSNP data for mouse, cat, chicken, cow, horse, pig, sheep, zebrafish and macaque.
- New phenotype data available for mouse (MGI, IMPC), rat (RGD), zebrafish (ZFIN), as well as updated OMIA and AnimalQTL data for several species.
Find out more
If you would like to learn more about these exciting updates, see live demos of the new features, and ask questions to the Ensembl team, please register for the release webinar at 4pm (GMT) on Wednesday 13th of December.
